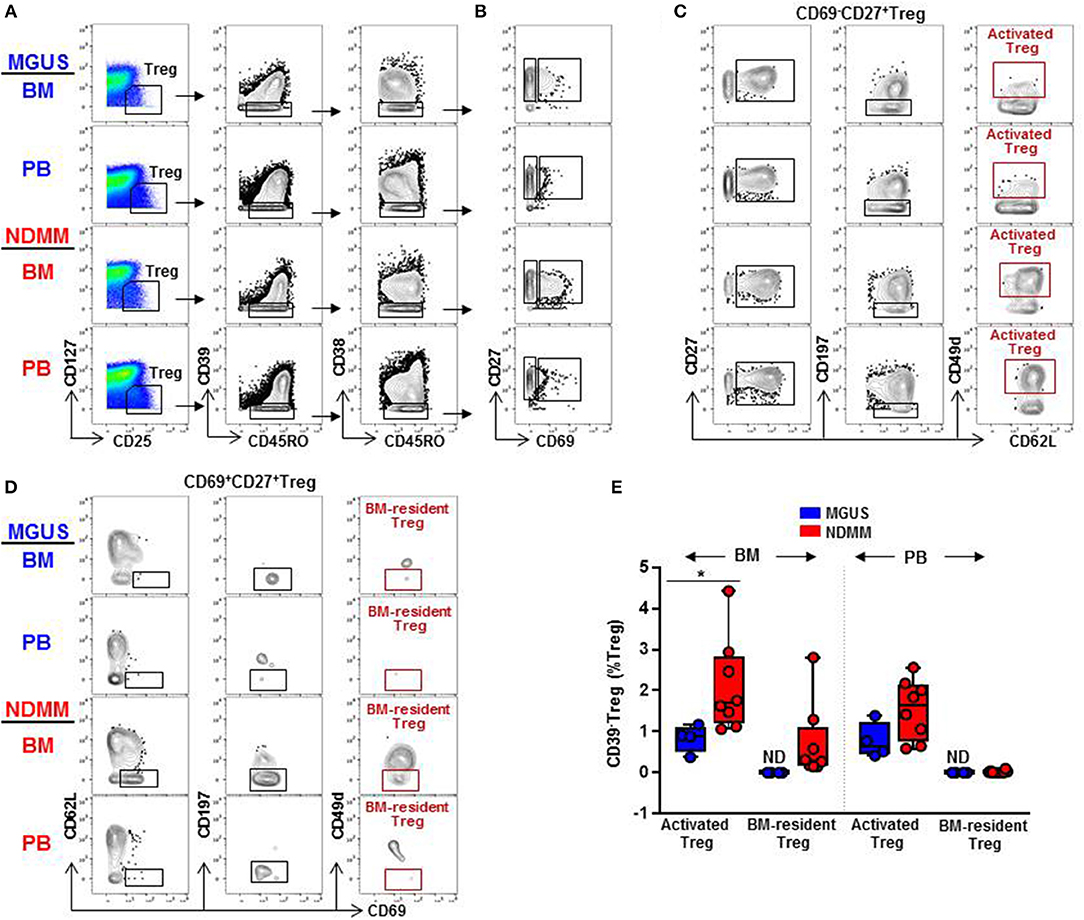
Frontiers | Mass Cytometry Discovers Two Discrete Subsets of CD39−Treg Which Discriminate MGUS From Multiple Myeloma
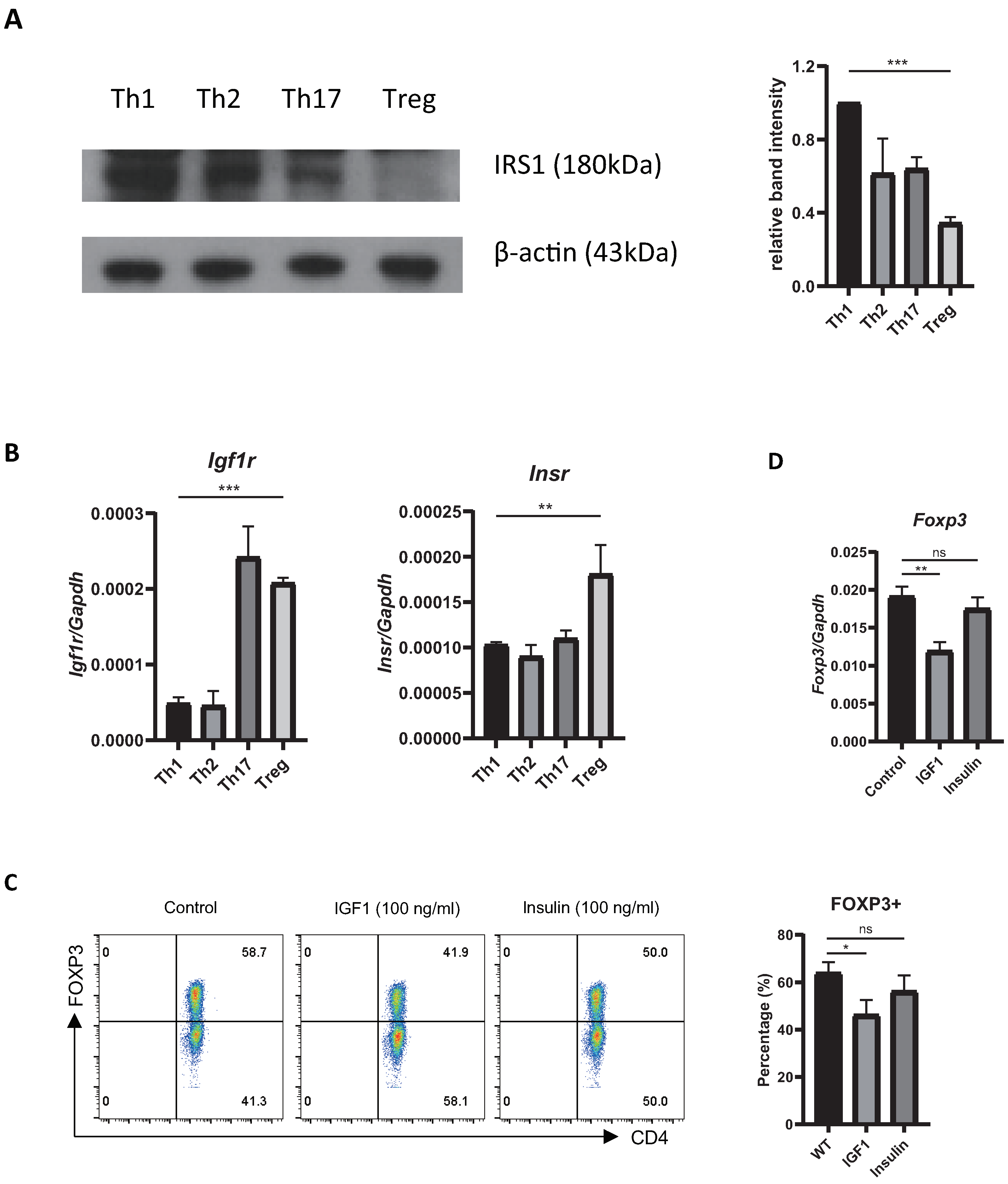
IJMS | Free Full-Text | Insulin Receptor Substrate 1 Signaling Inhibits Foxp3 Expression and Suppressive Functions in Treg Cells through the mTORC1 Pathway

Coya Therapeutics on X: "Coya Therapeutics to Present Data on Immune System and Regulatory #TCell (Treg) Contributions in #FrontotemporalDementia (#FTD) Patients at AD/PD 2024 Conference. Read the press release here: https://t.co/vXlwQMrrMb $COYA #

Treg analysis in rats. The percentage of CD4+CD25+Foxp3+ Tregs (a).... | Download Scientific Diagram

Expression of Tim-3 drives naïve Treg to an effector-like state with enhanced suppressive activity | bioRxiv
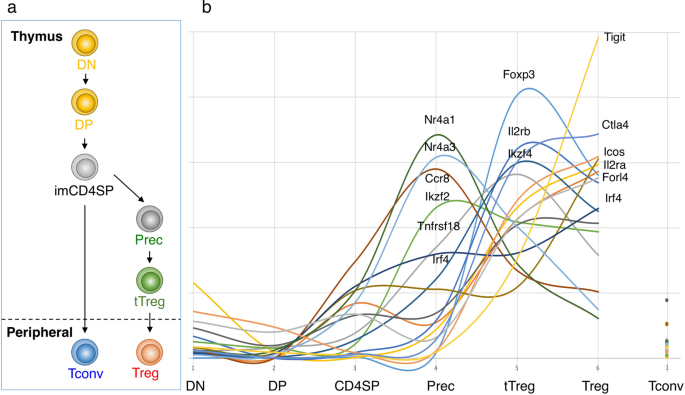
Transcriptional and epigenetic basis of Treg cell development and function: its genetic anomalies or variations in autoimmune diseases | Cell Research
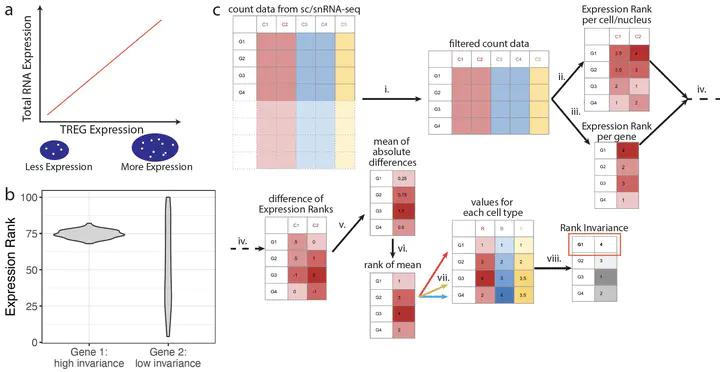
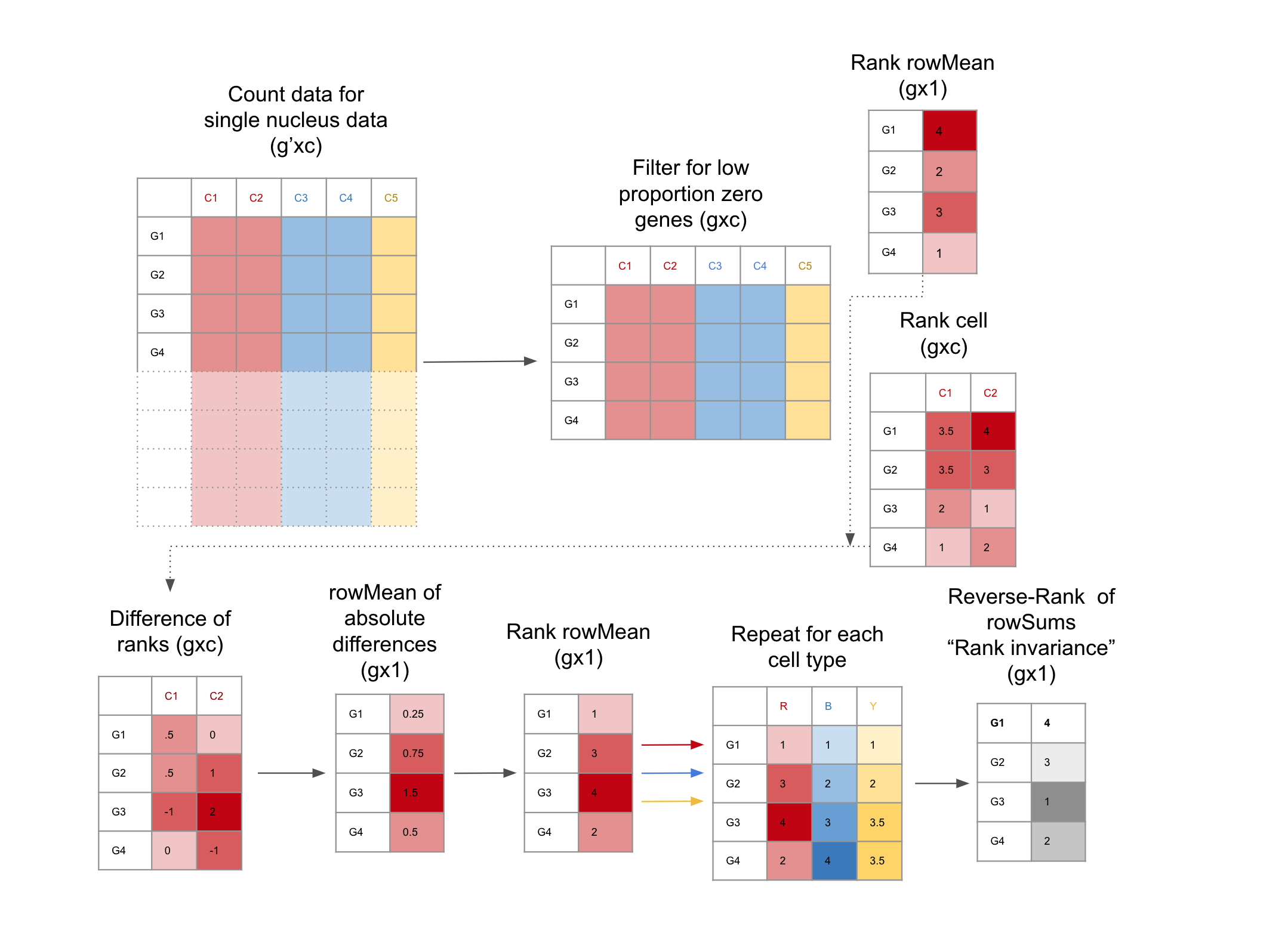
Data-driven Identification of Total RNA Expression Genes (TREGs) for Estimation of RNA Abundance in Heterogeneous Cell Types | L. Collado-Torres