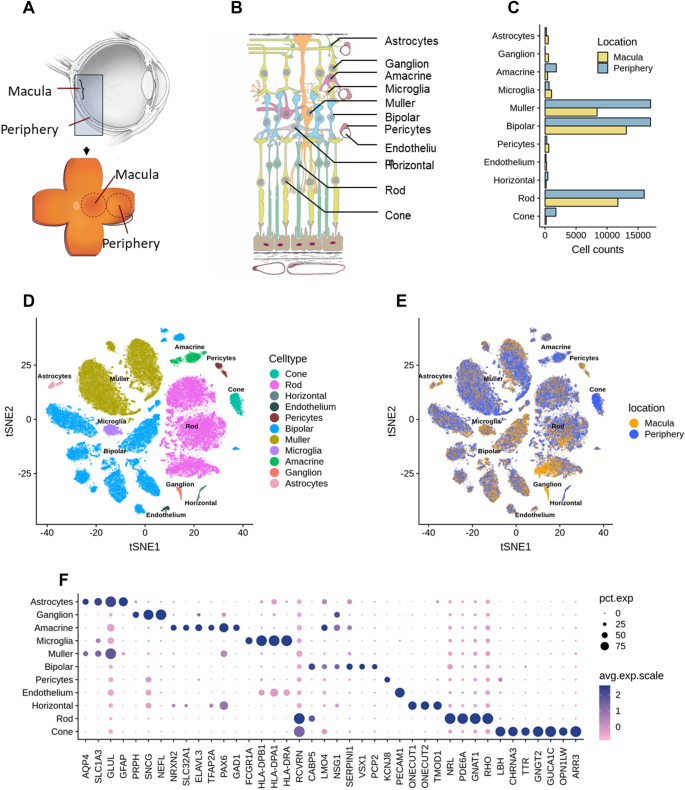
Implication of specific retinal cell-type involvement and gene expression changes in AMD progression using integrative analysis of single-cell and bulk RNA-seq profiling | Scientific Reports
Detecting cell-type-specific allelic expression imbalance by integrative analysis of bulk and single-cell RNA sequencing data | PLOS Genetics
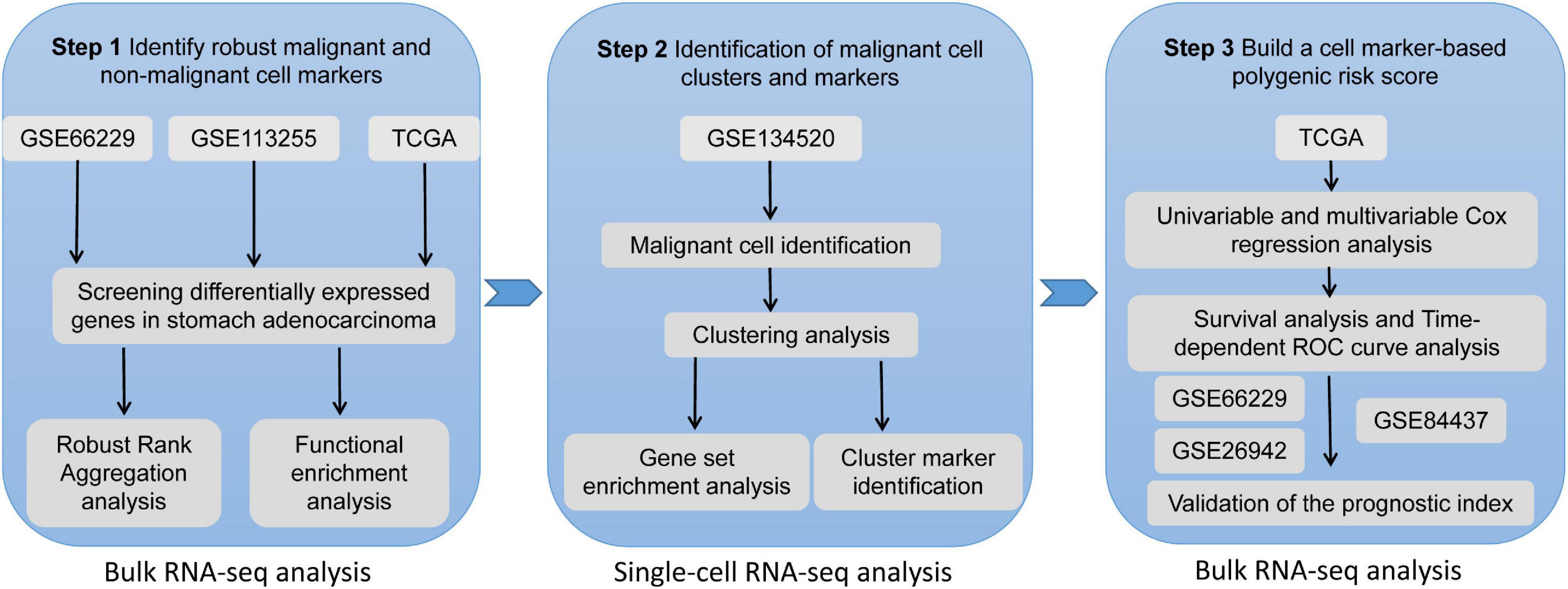
Frontiers | Identification and Validation of a Malignant Cell Subset Marker-Based Polygenic Risk Score in Stomach Adenocarcinoma Through Integrated Analysis of Bulk and Single-Cell RNA Sequencing Data
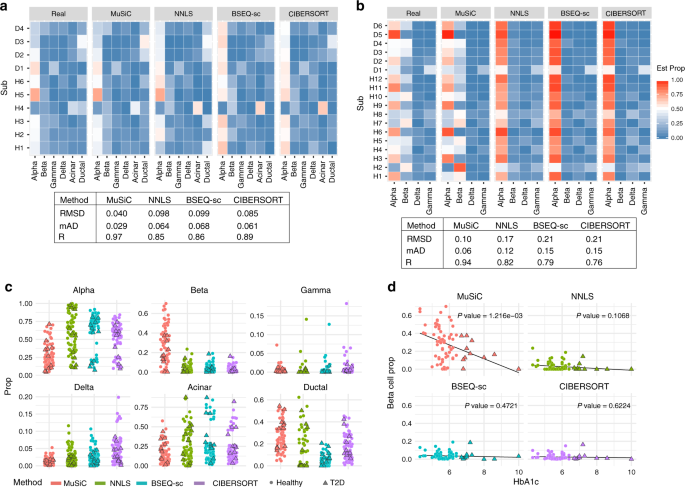
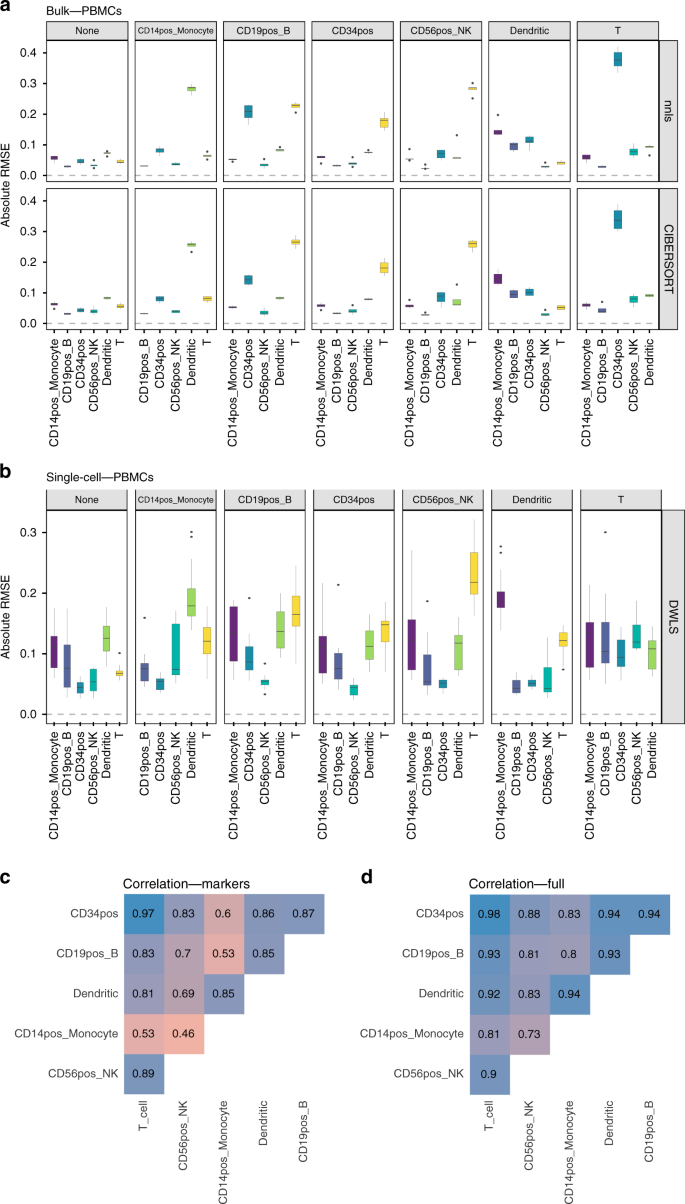
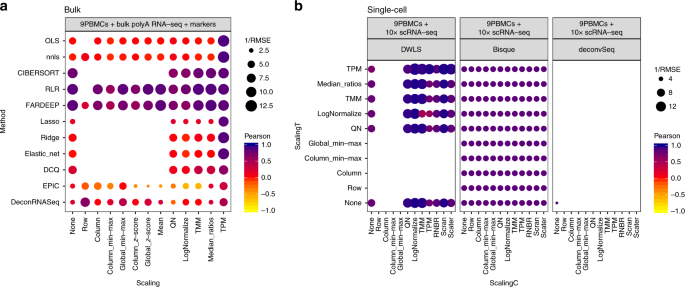
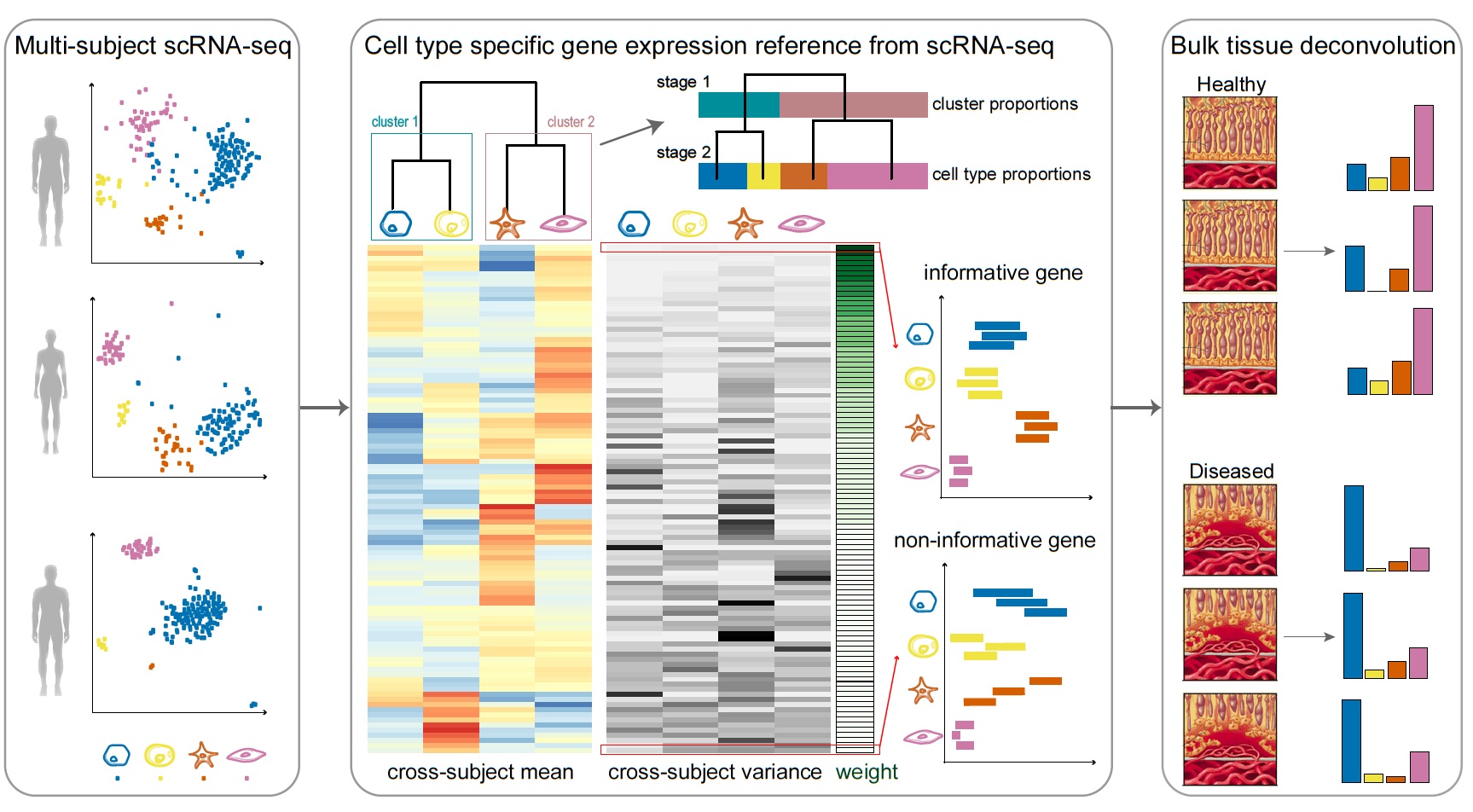
Bulk tissue cell type deconvolution with multi-subject single-cell expression reference | Nature Communications
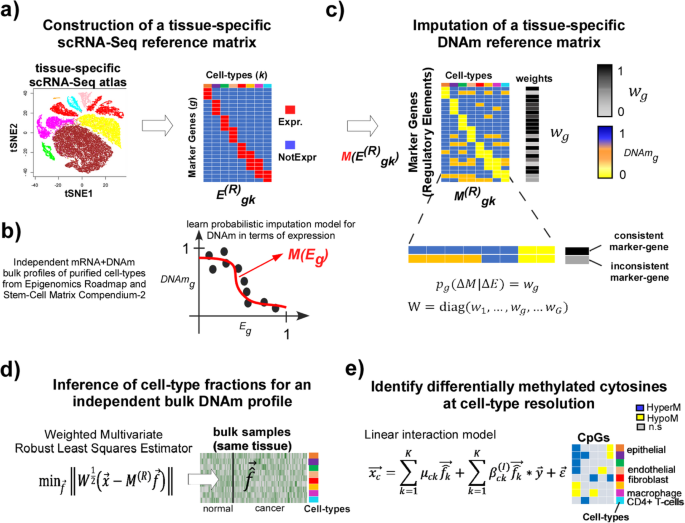
EPISCORE: cell type deconvolution of bulk tissue DNA methylomes from single- cell RNA-Seq data | Genome Biology | Full Text

A protocol to extract cell-type-specific signatures from differentially expressed genes in bulk-tissue RNA-seq: STAR Protocols

A protocol to extract cell-type-specific signatures from differentially expressed genes in bulk-tissue RNA-seq - ScienceDirect
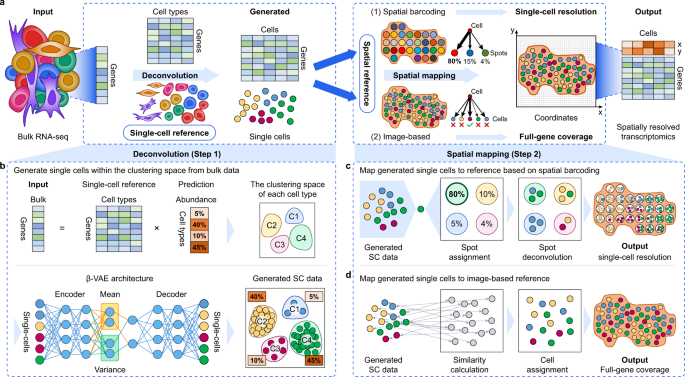
De novo analysis of bulk RNA-seq data at spatially resolved single-cell resolution | Nature Communications

Integrative single-cell and bulk RNA-seq analysis in human retina identified cell type-specific composition and gene expression changes for age-related macular degeneration | bioRxiv
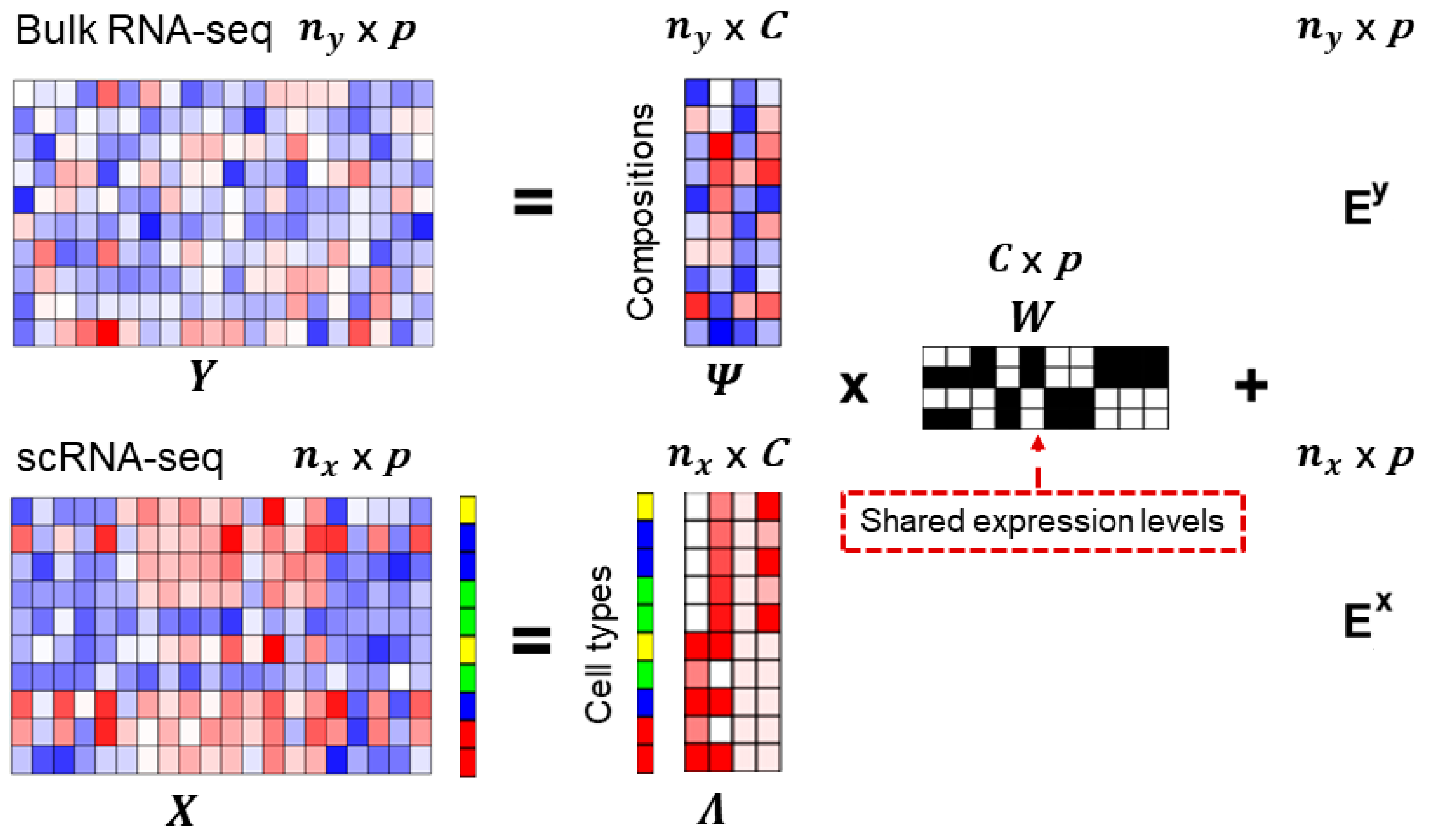
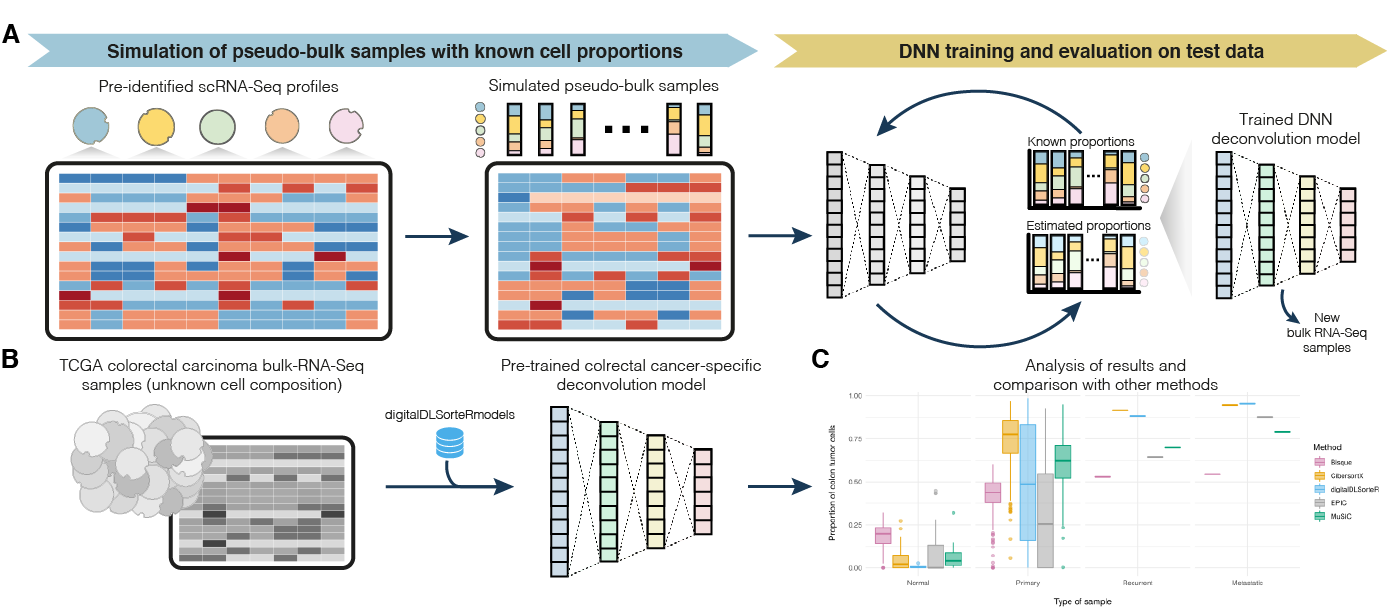
Cells | Free Full-Text | An Efficient and Flexible Method for Deconvoluting Bulk RNA-Seq Data with Single-Cell RNA-Seq Data
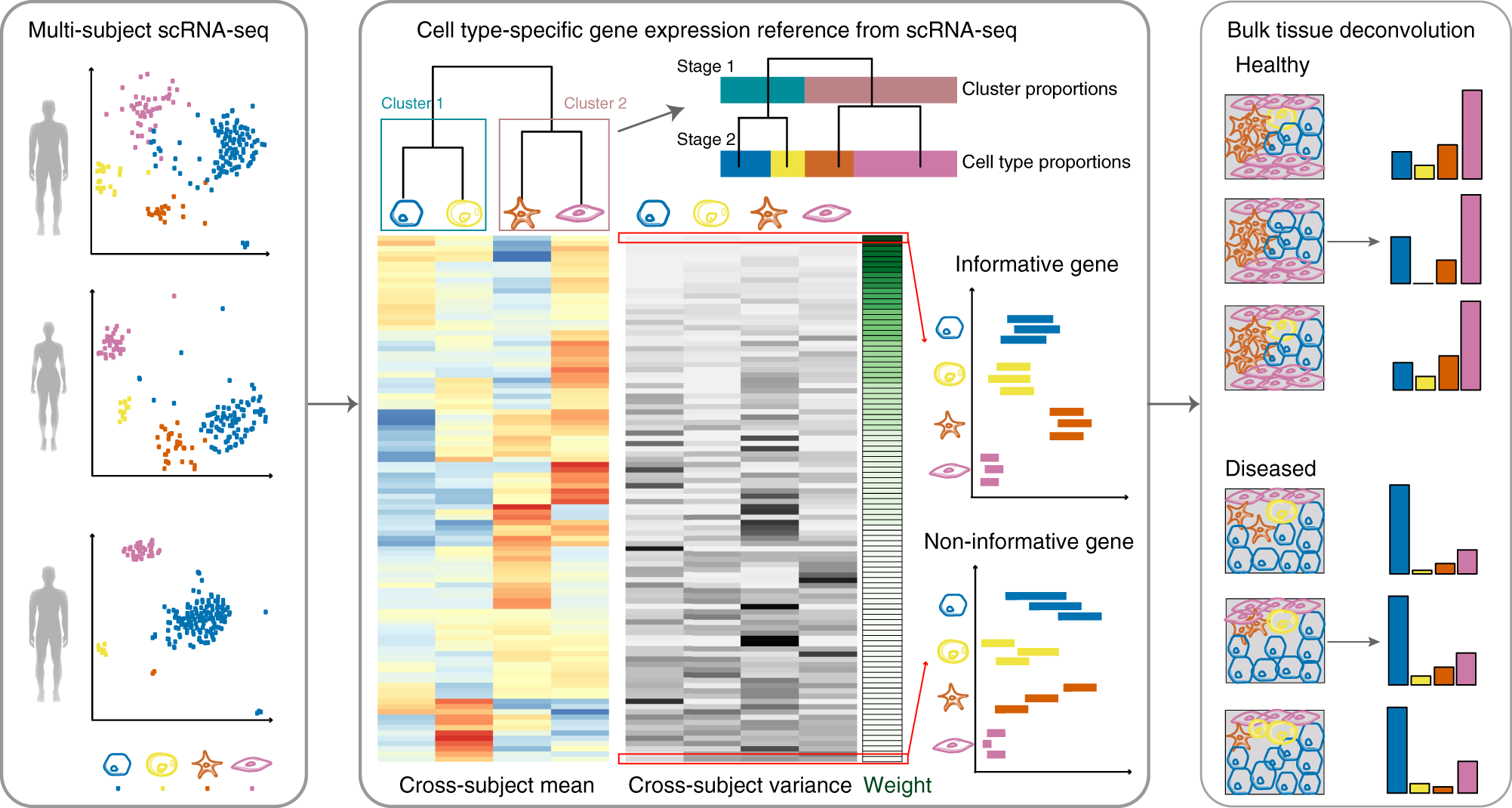
Bulk tissue cell type deconvolution with multi-subject single-cell expression reference | Nature Communications

SCDC – bulk gene expression deconvolution by multiple single-cell RNA sequencing references | RNA-Seq Blog
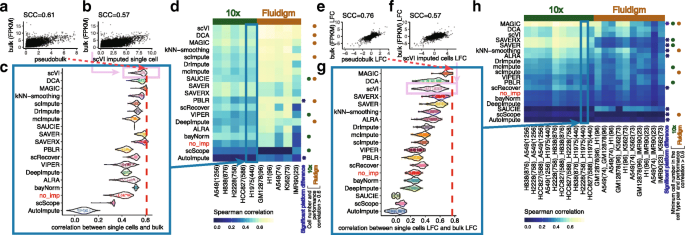
A systematic evaluation of single-cell RNA-sequencing imputation methods | Genome Biology | Full Text

![BingleSeq: a user-friendly R package for bulk and single-cell RNA-Seq data analysis [PeerJ] BingleSeq: a user-friendly R package for bulk and single-cell RNA-Seq data analysis [PeerJ]](https://dfzljdn9uc3pi.cloudfront.net/2020/10469/1/fig-1-2x.jpg)